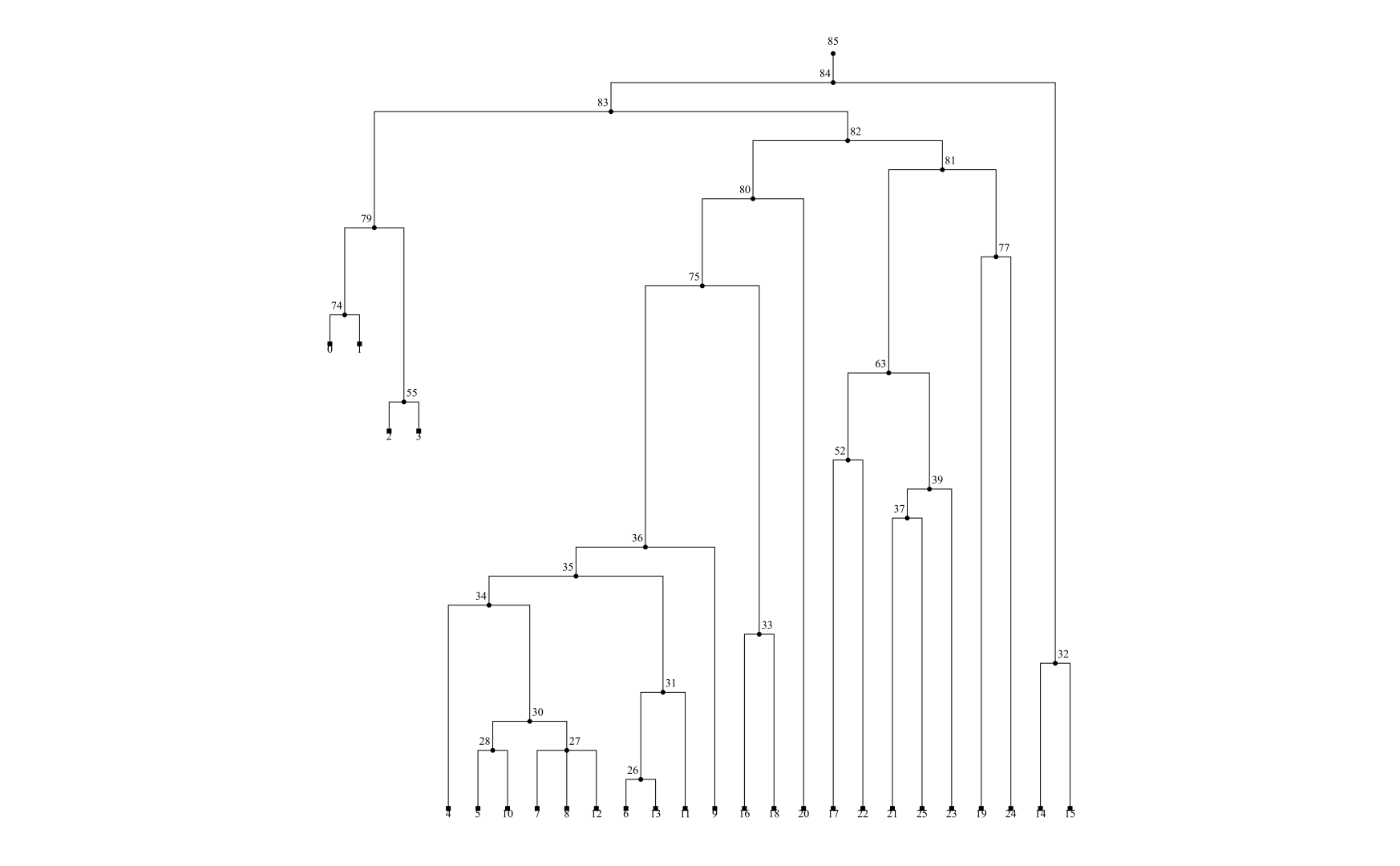
This function first obtains an SVG representation of the tree by calling the
draw_svg method of tskit and renders it as a bitmap image in R. All of
the many optional keyword arguments of the draw_svg method can be
provided and will be automatically passed to the method behind the scenes.
Usage
ts_draw(
x,
width = 1000,
height = 1000,
labels = FALSE,
sampled_only = TRUE,
title = NULL,
...
)Arguments
- x
A single tree extracted by
ts_tree- width, height
Pixel dimensions of the rendered bitmap
- labels
Label each node with the individual name?
- sampled_only
Should only individuals explicitly sampled through simplification be labeled? This is relevant in situations in which sampled individuals can themselves be among the ancestral nodes.
- title
Optional title for the figure
- ...
Keyword arguments to the tskit
draw_svgfunction.
Examples
check_dependencies(python = TRUE, quit = TRUE) # dependencies must be present
init_env()
#> The interface to all required Python modules has been activated.
# load an example model with an already simulated tree sequence
slendr_ts <- system.file("extdata/models/introgression_slim.trees", package = "slendr")
model <- read_model(path = system.file("extdata/models/introgression", package = "slendr"))
# load the tree-sequence object from disk
ts <- ts_load(slendr_ts, model)
# extract the first tree in the tree sequence and draw it
tree <- ts_tree(ts, i = 1)
# ts_draw accepts various optional arguments of tskit.Tree.draw_svg
ts_draw(tree, time_scale = "rank")